Over the past ten years, research on the process of adaptation and tolerance of plants during cold stress has been carried out. However, the underlying mechanisms are complex and not well understood. Furthermore, we contemplate whether this technology could be a viable alternative to the controversial genetically modified (GM) rice or it may also eventually embroil into a limbo.Ībstract: Previous studies have reported that low temperature (LT) constrains plant growth and restricts productivity in temperate regions. This technology could thus provide a much-needed fillip towards engineering TFs for generating Pi use efficient plants for sustainable agriculture. Here, we review the applications and challenges of this technique for genome editing of the TFs for deciphering the function and plausible interactions across them. Clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein 9 (Cas9) system has provided an attractive paradigm for editing genome in plants. This raised a pertinent question whether there are any likely interactions across these TFs. Since the functional characterization of PHR1 in 2001, several other TFs have now been identified in these model plants. Studies on MYB TF PHR1 in Arabidopsis thaliana (Arabidopsis) and its orthologs OsPHRs in Oryza sativa (rice) have provided empirical evidence of their significant roles in the maintenance of Pi homeostasis. Pi deficiency triggers an array of spatiotemporal adaptive responses including the differential regulation of several transcription factors (TFs). Availability of phosphate (Pi), the only assimilable P, is often suboptimal in rhizospheres. To summarize, to change the layout of networks generated by Metascape, we need to unbundle edges and then rebundle them.Abstract: Phosphorus (P), an essential macronutrient, is pivotal for growth and development of plants. This final plot is a significant improvement over Figure 4. At the end, bundle edges to produce more visually compact edge patterns for easier interpretation (right). Unbundle edges first and them move the red nodes (left). Once you are happy with the new locations of the nodes, simply use Layout > Bundle Edges > All Nodes and Edges to bundle the edges again (right). This will straighten all edges, then you can move the nodes around (the result is in Figure 6, left). To adjust Metascape networks, we first need to use the menu option Layout > Clear All Edge Bends. The example is Figure 5.C taken from this PubMed entry. Floppy bundled edges can lead to unreadable networks. Simply moving the nodes for the red cluster results in floppy edges circled in red. We sometimes see unaesthetic network plots due to this limitation, Figure 5 is an example taken from a recent publication. Simply moving the nodes, for instance the red cluster in Figure 3, can result in floppy edges (Figure 4). Sometimes, users would like to rearrange the nodes in the exported network, in order to better illustrate its biological context. 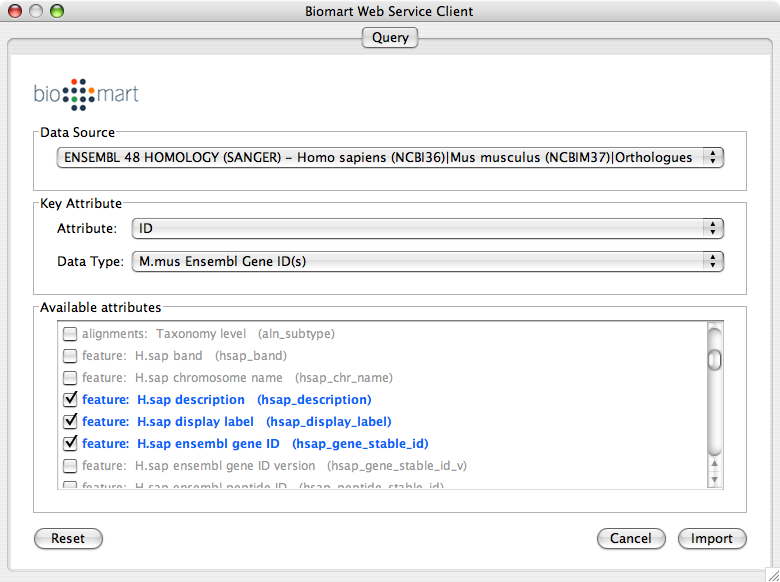
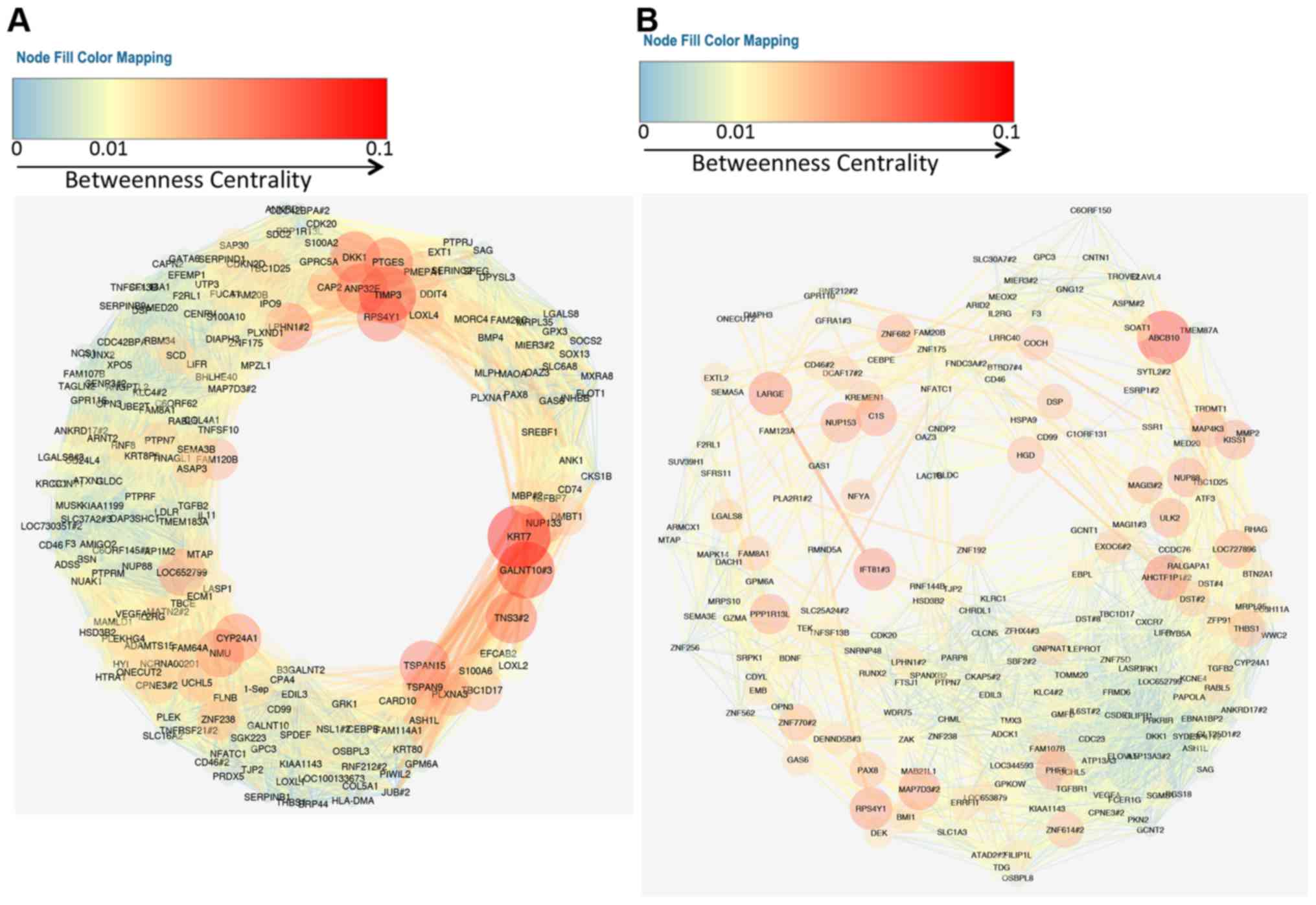
An enrichment network, where each node is colored by its cluster ID. For example, Figure 3 shows an enrichment network generated based on four input gene lists. Since edge bundling is so useful, it is used by default in network visualization outputs generated by Metascape. Operations lead to edge bundling in Cytoscape. The default parameters work well for most networks. To bundle edges in Cytoscape, use menu Layout > Bundle Edges > All Nodes and Edges (Figure 2). Screenshots were taken from a YouTube video here. High-level network edge patterns are more readily visible in the latter case. An example network rendered with straight edges (left) and bundled edges (right). Such visual clutter can be significantly reduced by an edge bundling algorithm implemented in Cytoscape (Figure 1). When a network contains too many edges, it can become a visual “hair ball” and no longer serves as an intuitive depiction. Metascape relies on Cytoscape to render networks, including both enrichment networks and protein-protein interaction networks.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
